SPROUT Classic
Design your own de novo ligands
SPROUT Classic is suitable for users who wish to generate small drug molecules by connecting up in-built as well as imported fragments.
SPROUT
Key features
A sophisticated tool for the design of drug-like, biologically active molecules.
An interactive system that can assist in several stages of the structure-based rational drug design process.
What’s in the box?
- Is a sophisticated de novo software tool
- Opens the crystal structure of target molecules in PDB format
- Has an advanced 3D graphical visualisation tool for displaying the detected target site regions, the generated structures, the protein receptor and the boundary surface
- Provides incredible control via
- Generates structures from fragments via new-bond, spiro and ring fusion joins
- Performs a systematic and exhaustive search to produce all the solutions that satisfy the steric constraints and the adjusted parameters
- Applies a bidirectional search to tackle the combinatorial explosion during the exploration of large search spaces
- Represents the search space and the target site regions continuously rather than as a set of discrete points
- Provides an option to perform the structure generation process remotely on a cluster by distributing the search among nodes
- Offers tools to help you select the structures of your interest based on
- Saves the results into PDB, MDL SDfile and Postscript formats
- Supports multi-session ligand design. This can be done by saving the results into SPROUT’s internal file format which can be reloaded at a later time in order to continue designing new structures
Modules in SPROUT
Binding pocket identification in the solvent accessible surface of the protein complex

Identification of favourable hydrogen bonding, metal bonding and hydrophobic regions or target sites within the binding pocket

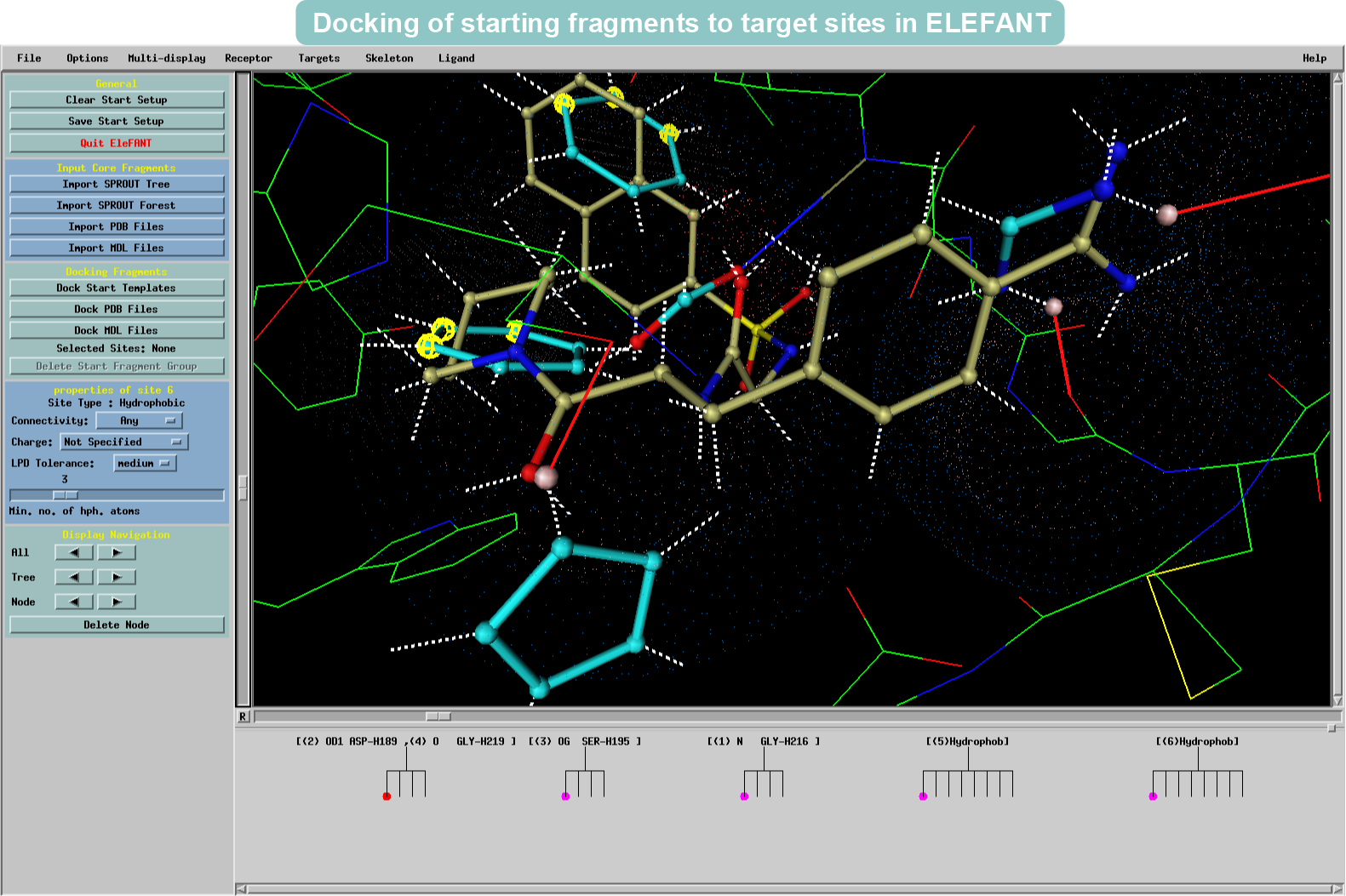
Docking of functional groups to target sites to form starting fragments for structure generation
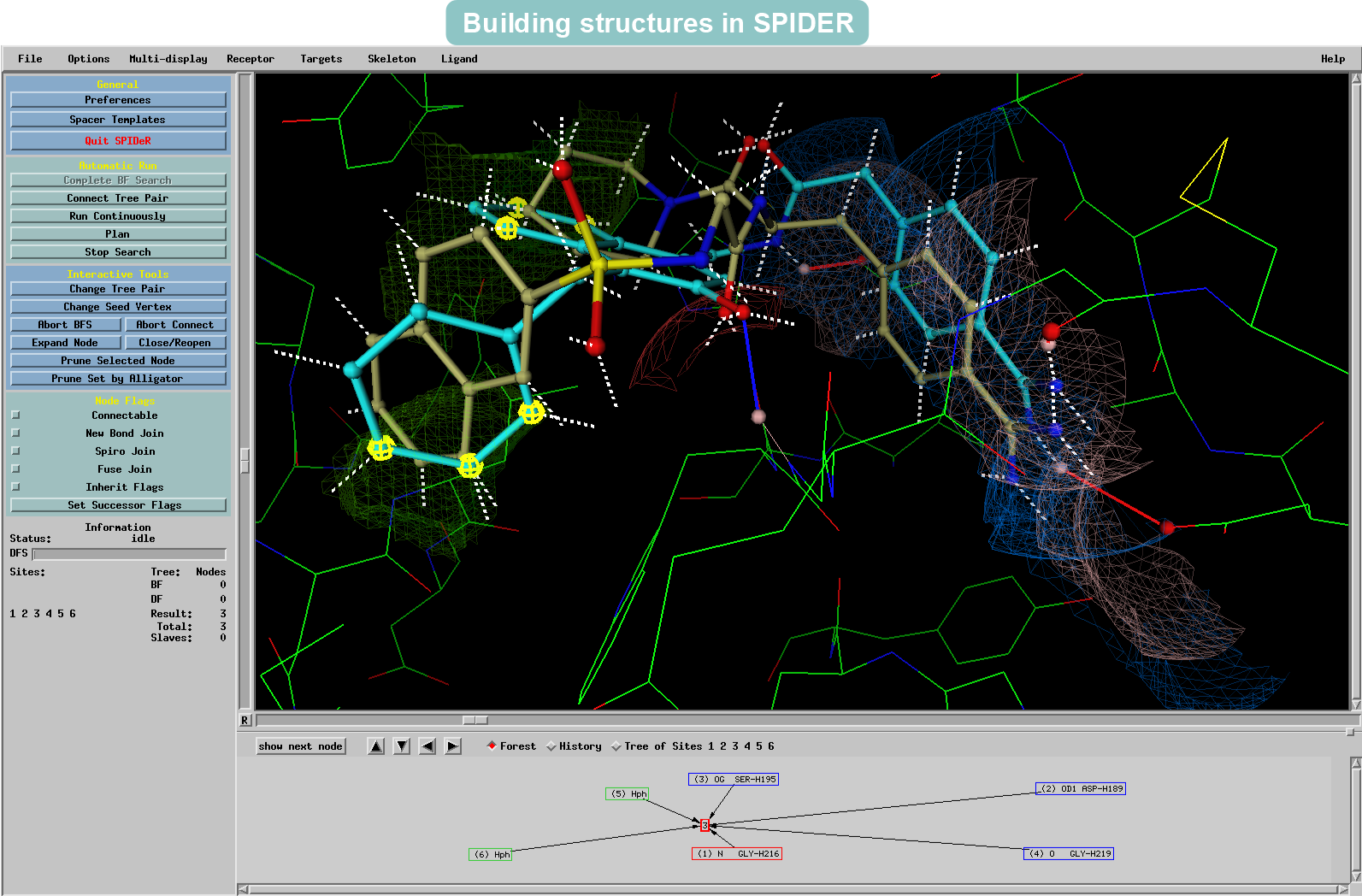
Fragment joining to build skeletons from the starting groups by satisfying the steric constraints of the binding pocket

Scoring and sorting solutions based on binding affinity, complexity analysis to estimate synthetic feasibility, and substructure searches
SPROUT in action
Thrombin has been identified as a key target for intervention in the blood coagulation cascade. Here SPROUT is used to generate structures that will have similarities with the NAPAP ligand bound to Thrombin. Using the original NAPAP ligand (displayed in the example as a brown skeleton) as the basis, new structures (displayed as light blue skeletons) are ensured to be tailored to suit the same cavity in the protein.
Get in touch
You need a high-performing, reliable and easy-to-use software solution to speed up your next big scientific breakthrough. Getting the right solution is integral to advance your research and workflow.